Header file for the GTF module. More...
#include <Python.h>#include <stdlib.h>#include "../classes/GtfDict/GtfDict.h"#include "../common.h"
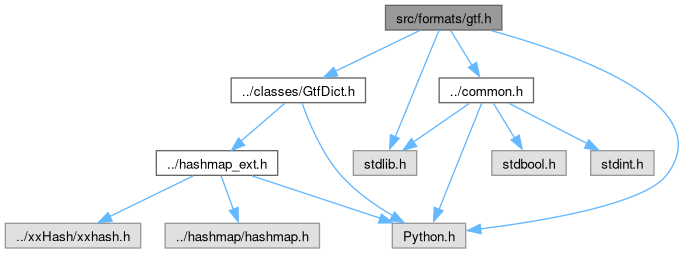
Include dependency graph for gtf.h:
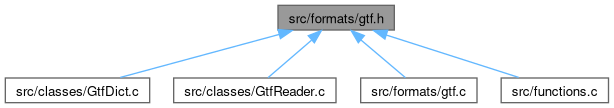
This graph shows which files directly or indirectly include this file:
Go to the source code of this file.
Macros | |
| #define | DEFAULT_ATTR_SIZE 64 |
| How many attributes should be allocated by default. | |
| #define | CORE_FIELD_COUNT 8 |
| The number of core fields in a GTF line. | |
| #define | GFF3_FASTA_HEADER "##FASTA" |
Enumerations | |
| enum | gtfFields { SEQNAME = 0 , SOURCE = 1 , FEATURE = 2 , START = 3 , END = 4 , SCORE = 5 , REVERSE = 6 , FRAME = 7 , ATTRIBUTES = 8 } |
| Enum containing the fields that a GTF line can contain. More... | |
Functions | |
| GtfDict * | createGTFdict (const occurrence_t *restrict token, PyObject *attr_tp, hashmap_t *restrict attr_keys, hashmap_t *restrict attr_vals) |
| Parses a singular GTF line into a Python dict. | |
| bool | validGTFLineToParse (const char *line, size_t len) |
| Determines if the provided line is a valid GTF line that can be parsed safely. | |
| char * | write_owned (char *restrict buf, size_t *restrict len, size_t *alloc, const char *restrict in, size_t inLen) |
| Writes a string to the output buffer, reallocing if necessary. | |
| char * | GtfDict_dump (GtfDict *restrict const self, char *restrict in, size_t *restrict len, size_t *restrict alloc) |
| Dumps the contents of a GtfDict to a string. | |
| int | GtfDict_as_annotation (PyObject *restrict object, long *restrict start, long *restrict end) |
| Converts a GtfDict or mapping object to an annotation. | |
Variables | |
| const char * | keywords [CORE_FIELD_COUNT] |
| Array containing the keywords of the GTF fields. | |
| const uint8_t | keyword_sizes [CORE_FIELD_COUNT] |
| Array containing the sizes of the keywords. | |
| const PyTypeObject * | keyword_types [CORE_FIELD_COUNT] |
| Array containing the types of the keywords. | |
Detailed Description
Header file for the GTF module.
Macro Definition Documentation
◆ GFF3_FASTA_HEADER
| #define GFF3_FASTA_HEADER "##FASTA" |
Function Documentation
◆ GtfDict_as_annotation()
| int GtfDict_as_annotation | ( | PyObject *restrict | object, |
| long *restrict | start, | ||
| long *restrict | end | ||
| ) |
Converts a GtfDict or mapping object to an annotation.
- Parameters
-
object the object to convert start a pointer to the start of the annotation end a pointer to the end of the annotation
- Returns
- 0 on success, -1 on failure
Here is the caller graph for this function:
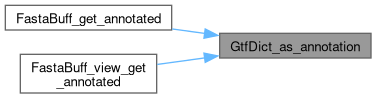
◆ GtfDict_dump()
| char * GtfDict_dump | ( | GtfDict *restrict const | self, |
| char *restrict | in, | ||
| size_t *restrict | len, | ||
| size_t *restrict | alloc | ||
| ) |
Dumps the contents of a GtfDict to a string.
- Parameters
-
self the GtfDict to dump in the input buffer to write to len a pointer to the length of the output string alloc a pointer to the size of the allocated output string
- Returns
- the output buffer
- Note
- if no allocation is done, in is returned unchanged
- Warning
- this function takes ownership of in and will free it if necessary
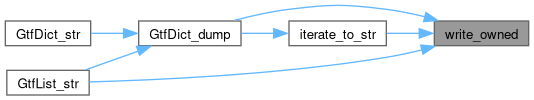
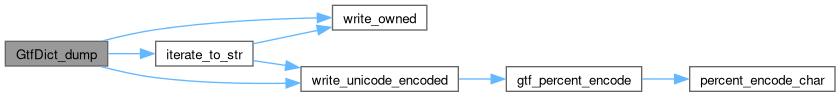
Here is the call graph for this function:
Here is the caller graph for this function:
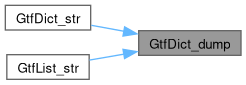
◆ write_owned()
| char * write_owned | ( | char *restrict | buf, |
| size_t *restrict | len, | ||
| size_t * | alloc, | ||
| const char *restrict | in, | ||
| size_t | inLen | ||
| ) |
Writes a string to the output buffer, reallocing if necessary.
- Parameters
-
buf The output buffer len The length of the output buffer alloc a pointer to the size of the allocated output buffer in The string to write inLen The length of the string to write
- Returns
- The updated output buffer
Here is the caller graph for this function: